Abstract
The objective of this study was to establish an accurate body weight (BW) prediction model for gilthead seabream Sparus aurata of 50–1000 g. Three thousand three hundred twelve (3312) fish were individually weighed and photographed. Traits measured from the images were total body length (TBL), fork body length (FBL), standard body length (SBL), body height (BH), head length (HL), eye diameter (ED), body area (BA, without fins), head area (HA), and eye area (EA). SBL, BH, BA, BA/SBL, and BA/BH showed a strong association with BW (correlation coefficients, r: 0.96–0.99). These traits were selected to proceed with the regression analysis. Simple, multiple linear, and 2nd-order polynomial regressions were applied to the whole data set and three BW subgroups of interest during gilthead seabream rearing (i.e., 50–100 g, 100–500 g, 500–1000 g). The prediction of BW from the whole data set was more accurate than from each BW subgroup. The models with the highest coefficient of determination (R2) and the lowest errors (mean absolute percentage error, MAPE) were either the power regression of BW with BA (R2: 99.0%, MAPE: 5.8%) or the multiple linear regression of BW with SBL, BA, BA/SBL, and BA/BH (R2: 98.6%, MAPE: 5.1%) as predictors. The accuracy of the two models is considered quite similar, and for reasons of simplicity, the power regression is advantageous, requiring only one trait to be measured (BA). The models identified in the present study can help to further develop the accuracy of machine vision-based systems for gilthead seabream BW measurement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Mediterranean marine intensive aquaculture has become one of the most important parts of the European Union primary production sector (HAPO 2023). This development is supported by continuous improvements in multiple aspects of the production cycle, e.g., fish handling and nutrition, fish health, and equipment (Føre et al. 2018; Li and Du 2022; Wu et al. 2022). A highly crucial point for the effective monitoring and management of the production progress is the knowledge, hopefully at any time, of fish body weight (BW) within sea cages (Tonachella et al. 2022; Chatziantoniou et al. 2023). Frequent evaluation of fish BW together with feed intake allows for estimation of fish growth performance and feed efficiency and thus for decisions regarding, e.g., grading, feeding, and harvest (Tonachella et al. 2022; Chatziantoniou et al. 2023).
In the aquaculture practice of marine Mediterranean fish species, the BW of the fish inside a cage is estimated through scheduled manual weighing of several small fish groups (Tonachella et al. 2022; Shi et al. 2023). The procedure requires labor effort, is time-consuming, and can be stressful to fish. Besides, there is the risk of error in the estimation since only a small sample of fish are weighed. Acknowledging the necessity for simplification and accuracy of the procedure, as well as for a “noninvasive, objective and repeatable measurement” (Li et al. 2020), several intelligent and automatic systems for BW estimation, mainly based on computer vision, have been developed, while research and interest on this field is continuously growing (e.g., Li et al. 2020; Li and Du 2022; Tonachella et al. 2022).
Not any kind of underwater equipment can weigh the fish, but instead, it can acquire images where fish morphometric traits can be visible and measurable. Thus, an important feature of any automatic weight-estimator system is the precise conversion of fish morphometric traits visible in the image to fish BW (Viazzi et al. 2015; Li et al. 2020; Holmes and Jeffres 2021; Li and Du 2022). The most common way to achieve the goal is to establish accurate regression models that predict fish BW from one or more traits. The weight-length relationship has been widely used for predictions of fish BW from total, standard, or fork length (e.g., gilthead seabream Sparus aurata, meagre Argyrosomus regius, red porgy Pagrus pagrus, Navarro et al. 2016; red tilapia Oreochromis niloticus, Jongjaraunsuk and Taparhudee 2022; gilthead seabream, Tonachella et al. 2022). However, available data suggest that the use of other morphometric data (e.g., surface area, body height) or the use of multiple traits can provide more accurate predictions (Alaskan Salmon species Oncorhynchus sp., Balaban et al. 2010a; Alaskan pollock Theragra chalcogramma, Balaban et al. 2010b; rainbow trout Oncorhynchus mykiss, Gümüş and Balaban 2010; herring Clupea harengus, Mathiassen et al. 2011; European seabass Dicentrarchus labrax larvae, de Verdal et al. 2014, Azevedo et al. 2023; Jade perch Scortum barcoo, Viazzi et al. 2015; Asian seabass Lates calcarifer, Konovalov et al. 2018, 2019; Nile tilapia, Fernandes et al. 2020; Taparhudee and Jongjaraunsuk 2023; European catfish Silurus glanis and African catfish Clarias gariepinus, Gümüş et al. 2021, chinook salmon Oncorhynchus tshawytscha, Holmes and Jeffres 2021; Australasian snapper Chrysophrys auratus, Yang et al. 2021; scaled and mirror carp Cyprinus carpio, Gümüş et al. 2023).
Given the importance of gilthead seabream Sparus aurata as a major reared species of intensive Mediterranean aquaculture, the objective of the present study was to establish the regression models needed to predict BW from traits measurable from images. Present data covered a BW range from 50 to 1000 g aiming to satisfy the needs of the main part of the production cycle in sea cages.
Materials and methods
Fish
A total of 3312 gilthead seabream were used in this study. Within the range of the targeted body weight (BW), i.e., 50–1000 g, special care was taken to have a minimum of 50 observations per 25 g weight classes (a total of 38 classes; in 3 out of 38 classes, the number of observations is 39–45 due to unsuccessful camera focus; see “Measurements”) (Table 1). The main bulk of samples (from 175 up to 1000 g) was obtained in a commercial packaging plant. In brief, harvested fish were slaughtered on site in 1 m3 bins with ice-seawater slurry and were thus transported to the packaging plant with refrigerator tracks. Upon arrival, fish were distributed according to their weight class (automatic sorting) and packaged (whole, ungutted, ventral side upwards), with flaked ice, in polystyrene boxes at 1–2 °C. Present measurements were performed in fish that had not reached full rigor mortis and not later than 12 h from slaughter. Fish of lower BW (from 50 up to 175 g) were obtained from a commercial fish farm. Although fish with BW ≤ 175 g were found among harvested fish in the packaging plant, they were not considered as representative of robust fish intensively growing. When the fish of a sea cage reach mean commercial weight and harvest begins, fish of much lower and much larger BW than the commercial size are also present, are harvested, and reach the packaging plant. However, the much smaller fish (i.e., <175 g) are either fish with low growth rates and/or fish that did not manage to feed well and thus were not suitable to be included in the present study. Instead, fish of 50–175 g, still growing in sea cages, are much more appropriate. On the farm, fish were netted from their cage and anesthetized in buckets. After weighing and photographing, fish recovered in buckets with clean sea water and were returned to their cage.
Measurements
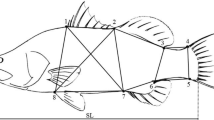
Each fish was individually weighed (scale precision 0.2 g for fish > 175 g and 0.1 g for fish <175 g), after wiping the surplus of water, and photographed. A photo camera (Canon, EOS M50) was mounted on a tripod, over the scale, and took lateral pictures of each fish. A ruler next to the fish was embedded in each photo to be used as a reference during morphometric traits measurements. The latter were performed using an image analyses software (Image-Pro Plus, v. 6.0). The software was set so that once the necessary landmarks were marked in the photo, the desired lengths (mm, Fig. 1), previously calibrated with reference to the ruler, were provided in an excel file. More precisely, the morphometric traits measured were total body length (TBL), fork body length (FBL), standard body length (SBL), body height (BH), head length (HL), and eye diameter (ED). Also, for each fish, the contour of its body (without fins), head, and eye was used to measure body area (BA), head area (HA), and eye area (EA) (one side, mm2).
Data analysis
All statistical analysis was carried out using Stata 18 software (Stata Corp, College Station, TX, USA). Data analysis was performed on the whole data set (i.e., 50–1000 g) and on three subgroups (Subgroup 1, S1: 50–100 g; Subgroup 2, S2: 100–500 g; Subgroup 3, S3: 500–1000 g). The weight range of the chosen subgroups corresponds to specific weight classes of importance to gilthead seabream intensive rearing. In particular, S1 has the fastest growing fish, while growth rate decreases as fish grow from S1 to S3; S2 includes fish that reach the commercial size, and S3 includes larger fish which are usually directed in special markets (e.g., restaurants), filleted, or selected as possible parents in breeding selection programs. The analysis of all the data together and in the three subgroups was decided in order to investigate whether more accurate BW prediction models can be obtained when the BW range is more limited.
For the whole data set and for each of the above-mentioned subgroups, Spearman rank correlations between BW and all morphometric traits measured, as well as the ratios BA/SBL, BA/BH, BA/HA, BA/EA, and SBL/BH, were calculated. Correlation coefficients were assessed according to Asuero et al. (2006) and Ratner (2009); special emphasis was given to identify a very high association of BW with a given morphometric trait (i.e., r>0.95). Based on correlation analysis results and taking into consideration that the fins may not be easily distinguished underwater while they contribute much to area but little to fish BW, regression models examined to predict BW were focused on SBL, BH, BA, BA/SBL, and BA/BH.
Simple, multiple linear, and 2nd-order polynomial regression models were applied to obtained data. In the case of simple regression, for all parameters examined, the power equation [Y = a Xb, Y: BW, X: morphometric traits or ratios, ln(a): intercept, b: slope] resulted in the highest coefficient of determination (R2) values, so other models are not included in the present study. In the case of multiple linear regression (Y = C0 + C1 X1 + C2 X2 + … + Ci Xi; Y: BW; C0 – Ci: constants; X1 – Xi: morphometric traits or ratios), stepwise regression with backward selection was applied. The analysis was performed including only the basic morphometric traits (i.e., SBL, BH, BA) or including only the ratios (i.e., BA/SBL, BA/BH) or including all data together (i.e., SBL, BH, BA, BA/SBL, BA/BH). In the tables, the final chosen model of each case is presented. Finally, each morphometric trait or ratio was also fitted to a 2nd-order polynomial regression model (Y = C0 + C1 X + C2 X2; Y: BW; C0 – C2: constants; X: morphometric traits or ratios). When the P value of the 2nd-order term of the polynomial was greater than 0.05, then the term was removed, and the order of the model was reduced to one.
To compare and assess the precision of the models obtained, the following metrics were used (Asuero et al. 2006; Ratner 2009; Chicco et al. 2021): (a) coefficient of determination (R2), as a measure of the proportion of the variance in the dependent variable (i.e., BW) that is predictable from the independent variables (i.e., morphometric traits). Models with R2 ≥ 98% were considered strong. (b) Mean absolute error (MAE), as an estimate of the absolute difference between the actual and predicted values. (c) Mean absolute percentage error (MAPE), as an estimate of the percentage difference between predicted values and actual values. MAPE is the percentage equivalent of MAE. (d) Root mean square error (RMSE), as a measure of the standard deviation of residuals. In the present data, the best models were initially sought among those with the highest R2 (Chicco et al. 2021). Once these were identified, then selection was refined among those models with the lowest error terms.
Results
Whole data set (50–1000 g)
The mean, median, standard deviation, minimum-maximum value, and coefficient of variation for BW and morphometric traits recorded and calculated (i.e., ratios) for the whole data set (BW range 50–1000 g) are shown in Table 2. Spearman rank correlation coefficients (Table 3) of BW with each of the traits determined were highly significant. Regarding the traits measured, the strongest correlation (r = 0.9904) was detected between BW and BA (monolateral, fins excluded), while the weakest was between BW and parameters related to head and eye (i.e., HL, HA, ED, EA). Correlations of BW with body lengths (i.e., TBL, FBL, SBL, BH) were also strong (r > 0.98). Regarding the ratios calculated, a strong correlation was found between BW and the ratios BA/SBL and BA/BH. Correlation coefficients for BW with BA/HA and BA/EA were lower, while no association is indicated between BW and the ratio SBL/BH.
Regression analysis performed in the present study focused on SBL, BH, BA, BA/SBL, and BA/BH as possible estimators of the BW (Table 4). Simple power regression (Eqs. 1–5, Table 4) showed that the relationship with the highest R2 and the lowest errors (MAPE, MAE, RMSE) was the one that considers BA as X (Eq. 3, Table 4, Fig. 2). Although the other power equations also had high R2, the error level was much higher. The investigation of multiple linear models (Eqs. 6–8, Table 4) pointed out the equation that involved SBL, BA, BA/SBL, and BA/BH (Eq. 8, Table 4). R2, as well as the level of errors observed, were comparable to values obtained in the power regression (Eq. 3, Table 4). Finally, the 2nd-order polynomial regression (Eqs. 9–13, Table 4) did not reveal a more accurate equation than those observed with power and multiple linear models. BA was the best estimator for BW (Eq. 11, Table 4), but still R2 and error levels were inferior to those observed for Eqs. 3 and 8.
Data subgroups (50–100 g, 100–500 g, 500–1000 g)
Descriptive statistics (Table SM1), correlation (Table SM2), and regression analysis data (Tables SM3-SM5, Fig. SM1-SM3) for each subgroup of BW examined are presented in Supplementary Material (SM). All correlation coefficients were slightly lower compared to those obtained when the whole data set was considered as one, but the results obtained regarding their relative importance were identical. Regression analysis applied to each subgroup showed, in all equations, lower R2 and similar or smaller error levels than those obtained from the whole data set. Apart from these differences, in each regression model category (i.e., power, multiple linear, 2nd-order polynomial), the more accurate (in terms of R2 and errors) equations obtained were the same as those detected for the weight range 50–1000 g, i.e., power equation with BA, multiple linear equation with the involvement of SBL, BH, and BA, and polynomial equation with BA, differing in the related constants.
Discussion
In the present study, morphometric traits considered to be suitable to predict BW in a regression model were those that were strongly (r > 0.95) correlated with BW. Strong (i.e., r > 0.80) association of BW with BA, SBL, BH, and HL has also been previously reported for gilthead seabream (Boulton et al. 2011; Navarro et al. 2016) and for other species (i.e., de Verdal et al. 2014; Fernandes et al. 2020). Similarly to the present results, a weak association was detected between BA and ED in spotted scat Scatophagus argus (Chen et al. 2022) and between BA and the ratio BH/SBL (the inverse of the present examined ratio) in pirapitinga Piaractus brachypomus (Ribeiro et al. 2019). Thus, all traits detected to be strongly associated with BW had adequate potential to be included in the regression models investigated, while traits like HL, HA, ED, EA, BA/HA, BA/EA, and SBL/BH could be excluded. Among body lengths, TBL and FBL were also excluded, despite the high correlation coefficients observed, since they involve the caudal fin which may lead to inaccurate measurements (Viazzi et al. 2015; Konovalov et al. 2019; Jongjaraunsuk and Taparhudee 2021).
The present study showed that models predicting BW of gilthead seabream from morphometric traits when the whole BW range is included as one data set are more accurate, in terms of R2, compared to those obtained when the defined subgroups of BW are used. On the other hand, models identified in the latter case had similar or lower values of the error terms. However, all MAPE (mean absolute percentage error) values obtained for the best equations identified in each regression model (no matter the data set) lay at the lower limit of percentage errors (4–14 %) reported in similar studies for other fish species (e.g., Viazzi et al. 2015, Konovalov et al. 2018, 2019; Fernandes et al. 2020, Jongjaraunsuk and Taparhudee 2021). Besides, according to Chicco et al. (2021), the coefficient of determination (R2) is considered as a more “informative and truthful” metric for the evaluation of regression analyses and for the comparison among different regression models. Overall, the present results suggest that the prediction of BW from the regression models of each BW subgroup examined does not offer an advantage over the prediction resulting by using the equations of the whole data set.
A common feature of all best BW prediction models obtained in the present study, no matter the BW range that they were applied, is the involvement of fish body area (BA) as an important predictor. Previously reported studies that investigated the best estimator variables for BW prediction and included fish area along with other measurements (e.g., contour, length, width, height, eccentricity) concluded that just fish area is sufficient to produce the more accurate regression models, at least for the species they studied (Balaban et al. 2010a; de Verdal et al. 2014; Viazzi et al. 2015; Fernandes et al. 2020; Gümüş et al. 2021; Taparhudee and Jongjaraunsuk 2023). Furthermore, the fact that fins were excluded from the present BA measurement probably adds to the accuracy of obtained models, as it was observed in Jade perch (Viazzi et al. 2015) and Asian seabass (Konovalov et al. 2019; Jongjaraunsuk and Taparhudee 2021), although no effect was found for Alaskan pollock (Balaban et al. 2010b). Fins contribute much to the area but little to BW; they continuously fold and unfold in a swimming fish, while they may also be damaged. The measurement of BA without the fins is a more accurate variable and explains the improvement of related predictions, whereas it may better fit in computer vision systems.
Focusing on the prediction models obtained from the whole data set (i.e., 50–1000 g), the models with the highest R2 and the lowest errors were either the power regression of BW with BA or the multiple linear regression of BW with SBL, BA, BA/SBL, and BA/BH as predictors. In the latter case, besides BA, two more measurements are involved, i.e., SBL and BH. The accuracy of the two models is quite similar, and for reasons of simplicity, the power regression is advantageous. In the literature, the power curve has been extensively used to evaluate allometric relationships between fish BW and mostly body length in fisheries and marine biology studies (e.g., Sangun et al. 2007; Robinson et al. 2010; Mathiassen et al. 2011; Karachle and Stergiou 2012; Sinopoli et al. 2022). Morphometric traits have also been of use in the aquaculture science to identify shape-related differences between farmed and wild fish (e.g., Arechavala-Lopez et al. 2012; González et al. 2016; Fragkoulis et al. 2017) and skeletal deformities (Costa et al. 2013), as well as to differentiate between the sexes (Çoban et al. 2011).
In conclusion, the power equation between BW and BA reported in the present study [ln(BW) = −8.846 +1.525 × ln(BA) or BW = exp (−8.846 + 1.525 × BA)] can accurately predict gilthead seabream BW, in the range of 50–1000 g, through the measurement of only one trait, i.e., monolateral body area without fins. It remains for the machine vision-based methods (Li et al. 2020; Li and Du 2022) to develop a way to acquire the image, extract the fins, and measure the body area by surpassing all the practical difficulties that an underwater measurement of a great number of swimming fish involves.
Data availability
Data will be available upon request.
References
Arechavala-Lopez P, Sanchez-Jerez P, Bayle-Sempere JT, Sfakianakis DG, Somarakis S (2012) Morphological differences between wild and farmed Mediterranean fish. Hydrobiologia 679:217–23. https://doi.org/10.1007/s10750-011-0886-y
Asuero AG, Sayago A, González AG (2006) The correlation coefficient: an overview. Crit Rev Anal Chem 36:41–59. https://doi.org/10.1080/10408340500526766
Azevedo A, Navarro LC, Cavalheri T, Santos HG, Martins I, Ozório R (2023) Morphological traits for allometric scaling of the European Sea Bass Dicentrarchus labrax (Linnaeus, 1758) from Southern Portugal population. J Fish Biol 103:425–438. https://doi.org/10.1111/jfb.15460
Balaban MO, Chombeau M, Cırban D, Gümüş B (2010) Prediction of the weight of Alaskan pollock using image analysis. J Food Sci 75:E552–E556. https://doi.org/10.1111/j.1750-3841.2010.01813.x
Balaban MO, Ünal Şengör GF, Soriano MG, Ruiz EG (2010) Using image analysis to predict the weight of Alaskan salmon of different species. J Food Sci 75:E157–E162. https://doi.org/10.1111/j.1750-3841.2010.01522.x
Boulton K, Massault C, Houston RD, de Koning DJ, Haley CS, Bovenhuis H, Batargias C, Canario AV, Kotoulas G, Tsigenopoulos CS (2011) QTL affecting morphometric traits and stress response in the gilthead seabream (Sparus aurata). Aquaculture 319:58–66. https://doi.org/10.1016/j.aquaculture.2011.06.044
Chatziantoniou A, Papandroulakis N, Stavrakidis-Zachou O, Spondylidis S, Taskaris S, Topouzelis K (2023) Aquasafe: a remote sensing, web-based platform for the support of precision fish farming. Appl Sci 13:6122. https://doi.org/10.3390/app13106122
Chen H, Li Z, Wang Y, Yang W, Shi H, Li S, Zhu C, Li G (2022) Relationship between body weight and morphological traits in female and male spotted scat (Scatophagus argus). Pak J Zool 54:1675–1683. https://doi.org/10.17582/journal.pjz/20210305150335
Chicco D, Warrens MJ, Jurman G (2021) The coefficient of determination R-squared is more informative than SMAPE, MAE, MAPE, MSE and RMSE in regression analysis evaluation. PeerJ Comput Sci 7:e623. https://doi.org/10.7717/peerj-cs.623
Çoban D, Yildirim Ş, Kamaci HO, Suzer C, Saka Ş, First K (2011) External morphology of European Seabass (Dicentrarchus labrax) related to sexual dimorphism. Turk J Zool 35:255–263. https://doi.org/10.3906/zoo-0810-3
Costa C, Antonucci F, Boglione C, Menesatti P, Vandeputte M, Chatain B (2013) Automated sorting for size, sex and skeletal anomalies of cultured seabass using external shape analysis. Aquac Eng 52:58–64. https://doi.org/10.1016/j.aquaeng.2012.09.001
de Verdal H, Vandeputte M, Pepey E, Vidal MO, Chatain B (2014) Individual growth monitoring of European sea bass larvae by image analysis and microsatellite genotyping. Aquaculture 434:470–475. https://doi.org/10.1016/j.aquaculture.2014.09.012
Fernandes AFA, Turra EM, de Alvarenga ÉR, Passafaro TL, Lopes FB, Alves GFO, Singh V, Rosa GJM (2020) Deep learning image segmentation for extraction of fish body measurements and prediction of body weight and carcass traits in Nile tilapia. Comput Electron Agric 170:105274. https://doi.org/10.1016/j.compag.2020.105274
Føre M, Frank K, Norton T, Svendsen E, Alfredsen JA, Dempster T, Eguiraun H, Watson W, Stahl A, Sunde LM, Schellewald C, Skøien KR, Alver MO, Berckmans D (2018) Precision fish farming: a new framework to improve production in aquaculture. Biosyst Eng 173:176–193. https://doi.org/10.1016/j.biosystemseng.2017.10.014
Fragkoulis S, Christou M, Karo R, Ritas C, Tzokas C, Batargias C, Koumoundouros G (2017) Scaling of body-shape quality in reared gilthead seabream Sparus aurata L. consumer preference assessment, wild standard and variability in reared phenotype. Aquac Res 48:2402–2410. https://doi.org/10.1111/are.13076
González MA, Rodriguez JM, Angón E, Martínez A, Garcia A, Peña F (2016) Characterization of morphological and meristic traits and their variations between two different populations (wild and cultured) of Cichlasoma festae, a species native to tropical Ecuadorian rivers. Arch Anim Breed 59:435–444. https://doi.org/10.5194/aab-59-435-2016
Gümüş B, Balaban MO (2010) Prediction of the weight of aquacultured rainbow trout (Oncorhynchus mykiss) by image analysis. J Aquat Food Prod Technol 19:227–237. https://doi.org/10.1080/10498850.2010.508869
Gümüş E, Yılayaz A, Kanyılmaz M, Gümüş B, Balaban MO (2021) Evaluation of body weight and color of cultured European catfish (Silurus glanis) and African catfish (Clarias gariepinus) using image analysis. Aquac Eng 93:102147. https://doi.org/10.1016/j.aquaeng.2021.102147
Gümüş B, Gümüş E, Balaban MO (2023) Image analysis to determine length-weight and area-weight relationships, and color differences in scaled carp and mirror carp grown in fiberglass and concrete tanks. Turk J Fish Aquat Sci 23:TRJFAS21260. https://doi.org/10.4194/TRJFAS21260
HAPO (2023) Aquaculture annual report 2023. Hellenic Aquaculture Producers Association (HAPO). Available in: https://fishfromgreece.com/wp-content/uploads/2023/10/HAPO_AR23_WEB-NEW.pdf. Accessed 28 Nov 2023
Holmes EJ, Jeffres CA (2021) Juvenile chinook salmon weight prediction using image-based morphometrics. North Am J Fish Manag 41:446–454. https://doi.org/10.1002/nafm.10533
Jongjaraunsuk R, Taparhudee W (2021) Weight estimation of Asian sea bass (Lates calcarifer) comparing whole body with and without fins using computer vision technique. Walailak J Sci Technol 18:9495. https://doi.org/10.48048/wjst.2021.9495
Jongjaraunsuk R, Taparhudee W (2022) Weight estimation model for red tilapia (Oreochromis niloticus Linn.) from images. Agr Nat Resour 56:215–224. https://doi.org/10.34044/j.anres.2021.56.1.20
Karachle PK, Stergiou KI (2012) Morphometrics and allometry in fishes. In: Wahl C (ed.) Morphometrics. InTech, pp. 65-86. http://www.intechopen.com/books/morphometrics/morphometrics-and-allometry-in-fishes. Accessed 28 Nov 2023
Konovalov DA, Saleh A, Domingos JA, White RD, Jerry DR (2018) Estimating mass of harvested Asian seabass Lates calcarifer from images. World J Eng Technol 6:15–23. https://doi.org/10.4236/wjet.2018.63B003
Konovalov DA, Saleh A, Efremova DB, Domingos JA, Jerry DR (2019) Automatic weight estimation of harvested fish from images. Digital Image Computing: Techniques and Applications (DICTA), Perth, WA, Australia, pp 1–7. https://doi.org/10.1109/DICTA47822.2019.8945971
Li D, Du L (2022) Recent advances of deep learning algorithms for aquacultural machine vision systems with emphasis on fish. Artif Intell Rev 55:4077–4116. https://doi.org/10.1007/s10462-021-10102-3
Li D, Hao Y, Duan Y (2020) Nonintrusive methods for biomass estimation in aquaculture with emphasis on fish: a review. Rev Aquac 12:1390–1411. https://doi.org/10.1111/raq.12388
Mathiassen JR, Misimi E, Toldnes B, Bondø M, Østvik SO (2011) High-speed weight estimation of whole herring (Clupea harengus) using 3D machine vision. J Food Sci 76:E458–E464. https://doi.org/10.1111/j.1750-3841.2011.02226.x
Navarro A, Lee-Montero I, Santana D, Henríquez P, Ferrer MA, Morales A, Soula M, Badilla R, Negrín-Báez D, Zamorano MJ, Afonso JM (2016) IMAFISH_ML: A fully-automated image analysis software for assessing fish morphometric traits on gilthead seabream (Sparus aurata L.), meagre (Argyrosomus regius) and red porgy (Pagrus pagrus). Comput Electron Agric 121:66–73. https://doi.org/10.1016/j.compag.2015.11.015
Ratner B (2009) The correlation coefficient: its values range between +1 / −1, or do they ? J Target Meas Anal Mark 17:139–142. https://doi.org/10.1057/jt.2009.5
Ribeiro FM, Lima M, da Costa PAT, Pereira DM, Carvalho TA, de Souza TV, Botelho HA, Silva FF, Costa AC (2019) Associations between morphometric variables and weight and yields carcass in Pirapitinga Piaractus brachypomus. Aquac Res 50:2004–2011. https://doi.org/10.1111/are.14099
Robinson LA, Greenstreet SPR, Reiss H, Callaway R, Craeymeersch JA, De Boois I, Degraer S, Ehrich S, Fraser HM, Goffin A, Jorgenson LL, Robertson MR, Lancaster J (2010) Length–weight relationships of 216 North Sea benthic invertebrates and fish. J Mar Biol Ass UK 90:95–104. https://doi.org/10.1017/S0025315409991408
Sangun L, Akamca E, Akar M (2007) Weight-length relationships for 39 fish species from the North-Eastern Mediterranean coast of Turkey. Turk J Fish Aquat Sci 7:37–40
Shi C, Zhao R, Liu C, Li D (2023) Underwater fish mass estimation using pattern matching based on binocular system. Aquac Eng 99(10):1016. https://doi.org/10.1016/j.aquaeng.2022.102285
Sinopoli M, Pipitone C, Badalamenti F, D’Anna G, Fiorentino F, Gristina M, Lauria V, Rizzo P, Milisenda G (2022) Effects of a trawling ban on the growth of young-of-the-year European hake, Merluccius merluccius in a Mediterranean fishing exclusion zone. Reg Stud Mar Sci 50:102151. https://doi.org/10.1016/j.rsma.2021.102151
Taparhudee W, Jongjaraunsuk R (2023) Weight estimation of Nile Tilapia (Oreochromis niloticus Linn.) using image analysis with and without fins and tail. J Fish Environ 47:19-32. https://li01.tci-thaijo.org/index.php/JFE/article/view/258089. Accessed 28 Nov 2023
Tonachella N, Martini A, Martinoli M, Pulcini D, Romano A, Capoccioni F (2022) An affordable and easy-to-use tool for automatic fish length and weight estimation in mariculture. Sci Rep 12:15642. https://doi.org/10.1038/s41598-022-19932-9
Viazzi S, Van Hoestenberghe S, Goddeeris B, Berckmans D (2015) Automatic mass estimation of Jade perch Scortum barcoo by computer vision. Aquac Eng 64:42–48. https://doi.org/10.1016/j.aquaeng.2014.11.003
Wu Y, Duan Y, Wei Y, An D, Liu J (2022) Application of intelligent and unmanned equipment in aquaculture: a review. Comput Electron Agric 199:107201. https://doi.org/10.1016/j.compag.2022.107201
Yang Y, Xue B, Jesson L, Wylie M, Zhang M, Wellenreuther M (2021) Deep convolutional neural networks for fish weight prediction from images. 36th International Conference on Image and Vision Computing New Zealand (IVCNZ), Tauranga, New Zealand, pp 1–6. https://doi.org/10.1109/IVCNZ54163.2021.9653412
Funding
Open access funding provided by HEAL-Link Greece. This work is part of the project entitled “Structured Light-Based fish BiOmass estimator - SaLTBOx” with grant number MIS 5033589, which was supported by the European Maritime and Fisheries Fund in the context of the Operational Programme “Maritime and Fisheries 2014-2020” and the Greek government.
Author information
Authors and Affiliations
Contributions
Nafsika Karakatsouli: Conceptualization, Methodology, Supervision, Formal analysis, Funding acquisition, Writing - Original Draft. Marina Mavrommati: Investigation, Data Curation, Formal analysis, Writing - Review & Editing, Eva Iris Karellou and Alexis Glaropoulos: Investigation, Data Curation, Resources. Alkisti Batzina: Investigation. Konstantinos Tzokas: Resources, Supervision, Writing - Review & Editing.
Corresponding author
Ethics declarations
Ethical approval
The study was carried out in accordance with the EU Directive 2010/63/EU, national laws (PD 160/91) for animal experiments, and the Bioethics Committee of the Department of Animal Science (Agricultural University of Athens). None of the fish died or suffered from injuries during the experiments.
Competing interests
The authors declare no competing interests.
Additional information
Handling Editor: Gavin Burnell
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karakatsouli, N., Mavrommati, M., Karellou, E.I. et al. Weight prediction of intensively reared gilthead seabream Sparus aurata from morphometric traits measured in images. Aquacult Int (2023). https://doi.org/10.1007/s10499-023-01343-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10499-023-01343-w